Missing places and ability to convert to hg38
For the four BC PRS loci, no variants were listed at the specified genomic position in gnomAD version 2.1.1, namely rs572022984, rs113778879, rs73754909, and rs79461387. gnomAD v3.1.2 also reported no variants for three of these four loci for the corresponding loci in hg38 defined by dbSNP. [23] (Supplementary Table 2). The rs572022984 locus was listed but with a total allele count of zero in the NFE samples (Table 2).
For two loci, the mutation to hg38 resulted in a change in alleles, i.e. for rs143384623 (hg19: 1-145604302-C-CT; hg38: 1-145830798-C-CA) and rs5500057:439-69hg (hg19: 1-145604302-C-CT; hg38: 1-145830798-C-CA). -145604302). : 9-133271182-TC). For rs143384623, the alternative allele change from CT to CA did not result in a significant change in AFs observed in gnomAD NFE samples (5142/13304 (0.39) in v2.1.1 vs. 24316/64610 (0.312) in v3. Fisher exact test side P= 0.14). For rs550057, the observed AFs appeared exactly opposite, namely 3786/14828 (0.26) for the T allele in gnomAD v2.1.1 and 49878/67552 (0.74) for the C allele in gnomAD v3.1.2. Therefore, 1-49878/67552 was considered as AF by gnomAD v3.1.2 at this biallelic site.
Allelic frequencies and technical artifacts reported in gnomAD version 3.1.2
For 39 of the 320 PRS loci listed with AF > 0 in gnomAD version 3.1.2, at least one observation of technical artifacts was reported: 38 loci as in low-complexity regions, 3 loci as localized in low signal quality. have been placed site, and 1 failed the allele-specific VQSR filter (Supplementary Table 2).
According to an absolute difference threshold of 0.016 (Supplementary Fig. 1), 24 loci were determined to be indicative of AF deviation compared to CanRisk (Fig. 1, Table 2). Absolute differences ranged from 0.03 to 0.71, and for 21 of these 24 loci (87.5%), technical artifacts were reported in gnomAD v3.1.2.
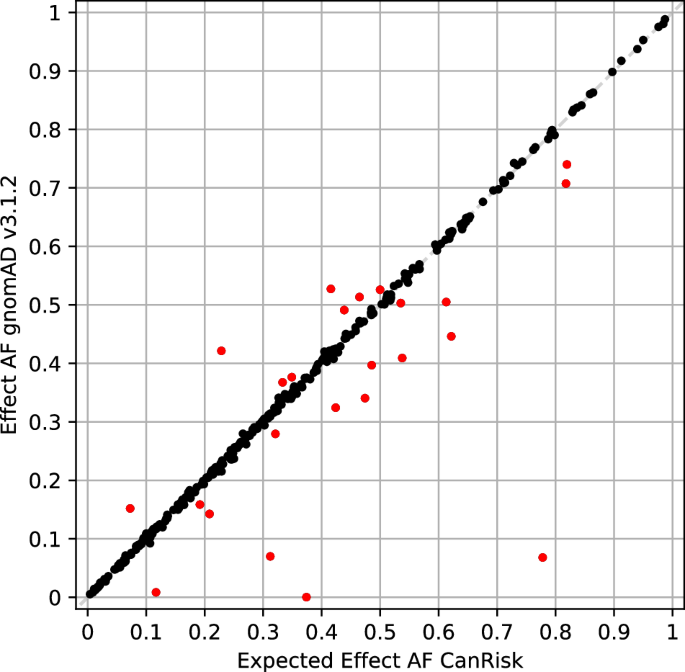
AF with severe deviation with absolute difference > 0.016 is indicated by red markers.
Evaluation of next-generation sequencing results in the real world
All 49 PRS sites for which significant AF deviation was observed in at least one of the datasets provided by the five GC-HBOC participating centers are listed in Table 3.
For the IMGAG DRAGEN data, 0.052 was calculated as the threshold for determining AFs with significant deviation (Supplementary Fig. 2), resulting in 18 affected loci (Table 3, Fig. 2). Of these, 16 were previously identified in gnomAD version 3.1.2 as missing or having significant AF deviation. The exceptions were rs62485509 and rs9931038. For the IMGAG freebays data, 0.036 was calculated as the threshold (Supplementary Fig. 2), resulting in 16 loci of BCAC 313 BC PRS indicating significant AF deviation. Of these, 11 loci were also identified as showing aberrant AF in the IMGAG DRAGEN data, and all but rs12406858 and rs11268668 were previously identified as missing or showing aberrant AF in gnomAD v3.1.2.
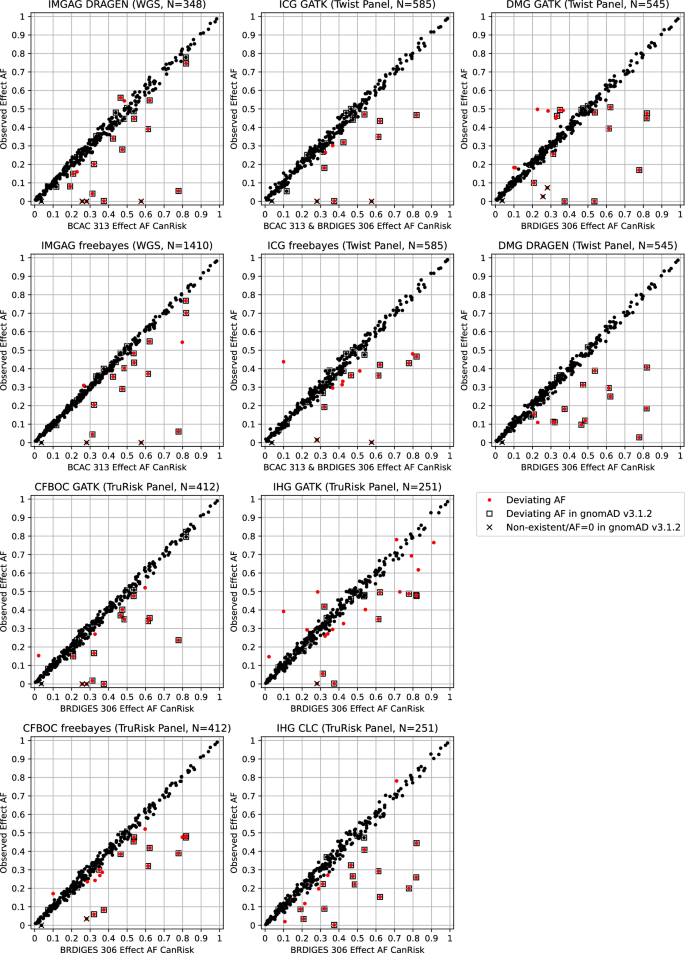
Data were provided by the Institute of Medical Genetics and Applied Genomics (IMGAG) at the University Hospital Tübingen, the Institute of Clinical Genetics (ICG) at the University Hospital Carl Gustav Carus Dresden, by the Department of Medical Genetics (DMG) at the University Hospital Münster, by the Familial Breast and Ovarian Cancer Center (CFBOC) at the University Hospital of Cologne, and by the Institute of Human Genetics (IHG) at the University of Regensburg.
Considering the genotype data provided by ICG based on 585 samples, 23 out of a total of 324 PRS loci did not meet the minimum quality criteria (read depth ≥ 20) in more than 25% of the samples and were discarded (Supplementary Table 3 ). In addition, GATK read depth <20 را برای > 25% of samples reported for rs56097627 and rs143384623. For 260 of the remaining 299 PRS loci (86.96%), forced genotyping with GATK and freebays resulted in the observation of identical AFs. For both GATK and freebayes ICG data, 0.063 was calculated as the threshold to determine significant AF deviation (Supplementary Fig. 3). Using this threshold, 11 AF sites showed significant deviation in the GATK dataset (including two sites unique to BCAC 313 BC PRS) and 14 sites in the freebayes dataset (including three sites unique to BCAC 313 BC PRS ) showed, with an overlap of 7 (Table 3, Figure 2).
DMG provided GATK-based PRS and DRAGEN 306 BC PRS genotyping data from 545 samples. Locus rs138179519 did not meet the quality criteria and additionally rs774021038 using DRAGEN. Of the remaining 304 loci, 252 (82.89%) showed the same AF (Supplementary Table 3). Using a threshold of 0.052 (Supplementary Fig. 4), 20 loci showed AF deviation in the GATK data and 14 loci in the DRAGEN data, respectively, with an overlap of 9 loci.
For CFBOC data based on 412 samples, a threshold of 0.047 was calculated (Supplementary Figure 5). BRIDGES 306 BC PRS loci were considered, 243 (79.41%) of which showed the same AF for both callers (Supplementary Table 3). A total of 25 loci (all of which were also included in BCAC 313 BC PRS) showed aberrant AFs: 16 loci in GATK and 19 loci in freebayes data, with an overlap of 10 loci.
IHG provided GATK- and CLC-based PRS genotyping data of 306 BC PRS from 251 samples (Supplementary Methods). Four sites in both settings, and another four sites in the CLC setting, did not meet the quality criteria. Of the remaining 298 loci, 228 (76.51%) showed the same AF (Supplementary Table 3). Using a threshold of 0.063 (Supplementary Fig. 6), resulted in 23 loci showing significantly deviant AF in the GATK data, 19 loci in the CLC data, respectively, with an overlap of 10 loci.
Briefly, for four loci, aberrant AFs were investigated in all GC-HBOC real-world settings, namely for rs56097627, rs113778879, rs57589542, and rs3988353. Four other loci, namely rs574103382, rs73754909, rs3057314, and rs57920543, were reported to have AF bias in all but one setting (Table 3).
However, 16 loci were found exclusively in one setting, namely five loci in the IHG GATK data (rs1511243, rs4880038, rs1027113, rs12709163, rs1111207), three each in the ICG data (rs1504893, 349, 371, 9504893) in the IHG data. rs10975870, rs11049431, rs144767203), two in DMG GATK data (rs10644978, rs66987842) and one each in IMGAG DRAGEN (rs9931038), IMGAG5000, IMGAG120 702307). Three other loci (rs10074269, rs55941023, rs35054928) showed AF aberrations in only one center, but these were concordant.
Considering the absence of the locus in gnomAD v3.1.2, rs113778879 was not observed with the expected AF in any GC-HBOC center, and rs73754909 was not observed in the DMG data only by forced DRAGEN calling. For rs79461387, the expected autofocus was consistently reported when using freebayes, but not with non-forced DRAGEN calling and in two settings using forced GATK. Notably, rs572022984 with an allele count of zero in gnomAD v3.1.2 NFE and an expected AF of 0.0364 in CanRisk was consistently not observed at all or with a maximum AF of 0.0037 (Supplementary Table 3).
Five loci representing aberrant AF in gnomAD v3.1.2 NFE (Table 2) with AF deviation by none of the participating GC-HBOC centers, namely rs78425380, rs62331150, rs60954078, rs1929, rs1925 and rs108, rs6095, rs108, rs1925, rs1925, rs1925 , 108, 108 and 1978, 1978, 108, 1975, 1978 and 1985, 18425380, with AF deviation not reported.
Concepts in risk prediction
Without additional information and assuming a standardized PRS at the 50th percentile, the estimated 10-year risks of developing primary BC for 20-, 40-, and 60-year-old women with cancer were 0.1%, 1.5%, and 3.4%. CanRisk (Supplementary Table 4). The percentage of PRSs from synthetic VCF files with aberrant doses (see “Materials and Methods”) ranged from 47.5% (IHG CLC, BRIDGES 306) to 55.7% (ICG freebayes, BCAC 313). The 0.1% risk for a 20-year-old woman was unchanged in all scenarios including artificial PRS. For a 40-year-old woman, the estimated 10-year risks increased by 0.1% in seven scenarios and by 0.2% in eight scenarios for a 60-year-old woman.
Assuming the median PRS (50th percentile) of 20-, 40-, and 60-year-old women with cancer, the estimated lifetime residual risks of developing primary BC based on CanRisk (Supplementary Table 4) are 11.3%, 10.9%, and 7.1%. When using PRS from synthetic VCF files with abnormal doses, estimated lifetime risks ranged from 11.1% to 11.9% for a 20-year-old woman, from 10.6% to 11.4% for a 40-year-old woman, and from 7.0 % to 7.4% for a 60-year-old woman. The lowest estimates were obtained with BRIDGES 306 BC PRS based on IHG CLC data with 19 synthetic doses entered, and the highest with PRS BCAC 313 BC based on ICG freebays data with 14 synthetic doses.
Consideration of alleles and alternative loci in linkage disequilibrium
For the 20 PRS loci showing significantly skewed AF in at least one real-world NGS dataset, alternative alleles or overlapping variants with an AF of at least 0.01 in NFE were reported in gnomAD version 3.1.2 (Supplementary Table 5). For rs73754909 and rs79461387, both SNVs and absent in gnomAD version 3.1.2, deletions with AFs comparable to those expected by CanRisk were reported. For both deletions, the adjacent downstream nucleotide of the reference sequence was identical to the replacement nucleotide of the expected effect allele (Figure 3). For rs113778879, which is also a SNV not present in gnomAD v3.1.2, similar observations can be made (Supplementary Fig. 7), but the reported AF is greater than expected by more than 0.1 (0.5762 vs. 0.6818).
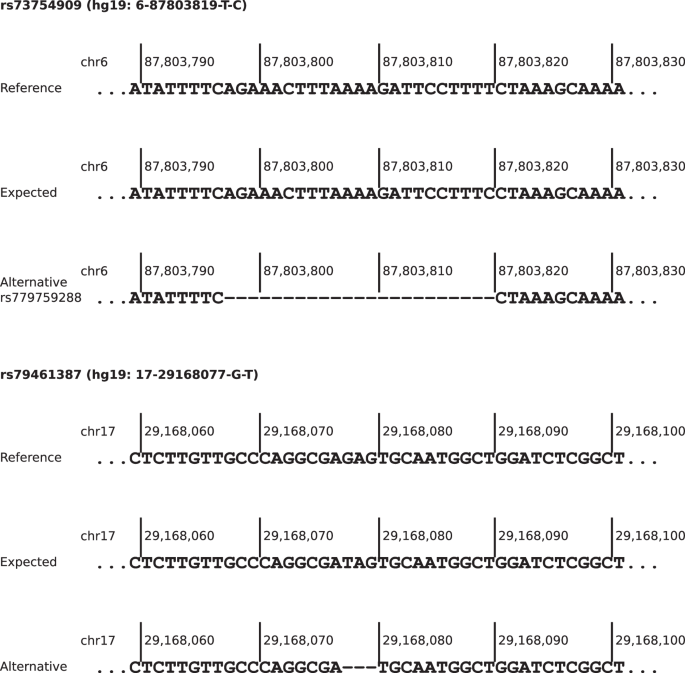
Both alternative alleles are deletions where the adjacent downstream nucleotide is identical to the expected alternative allele.
For 28 of the 49 loci showing significant AF deviation in at least one real data set, proxies could be identified in the 1000G GRCh37 microarray data, the 1000G GRCh38 high-coverage WGS data, or the European TOPMED data (Supplementary Table 6). For rs113778879, rs73754909, and rs79461387, LDpair based on GRCh38 reported the same alternative alleles as gnomAD v3.1.2 (Supplementary Table 5), where the major PRS loci are absent.
Proxies and alternative alleles showing AF in gnomAD v3.1.2 comparable to expected CanRisk AF, i.e., absolute deviation <0.016, were considered as possible solutions for improved PRS genotyping and further consideration of AF. observed in the IMGAG freebayes data were evaluated (Table 4). ). For 19 of these 21 PRS loci, the absolute difference between the expected and observed AF in the IMGAG freebayes data remained below the 0.036 threshold for IMGAG freebayes defined earlier. Exceptions were the rs12406858 and rs79461387 substitutions. The latter is noteworthy because the main PRS locus, which is a SNV, was correctly called by freebayes in both forced and unforced mode (Table 3), while GATK HaplotypeCaller seemed to remove GAG sequence overlap in the DMG data. and calls CFBOC. Also of note are the potential substitutions rs73754909 and rs111833376, as both variants are called AF variants with significant deviation in most real-world datasets.
#Limitations #nextgeneration #sequencingbased #genotyping #breast #cancer #polygenic #risk #loci #European #Journal #Human #Genetics